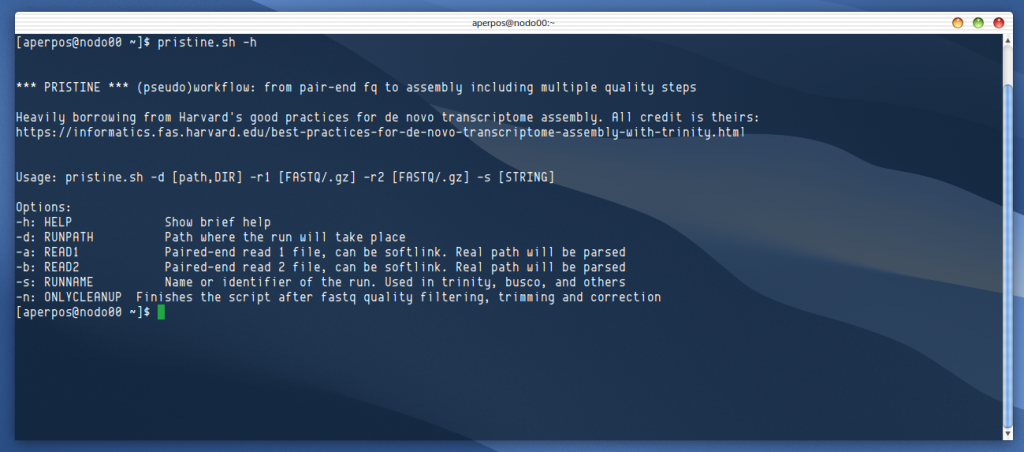
This is my first kind of serious project in bioinformatics. I had to prepare some de novo transcriptome assemblies from weird organisms using publicly available data, and I took the chance to learn a little bit how to automate processes using bash scripting, virtual environments, a a lot of variables and flags.
I have named this pristine, and it can be found in my github repository.
I will keep working on it as I learn how to code and make new things. I recently saw a way to download and transfer fastq data into other softwares on-the-go as it is downloaded using UNIX pipes. I will try to check if something like this could be done, how cool.
Cheers!
(and no, I did not forget about the last post of the multicellularity story. I just need free time and energies to sit down and finish it :’) )